It’s not just genomic scientists that are dealing with enormous amounts of DNA sequence data – the clinician will soon be next. However, the explosion in linking disease with genetics, and the realization that the FDA will require gene testing prior to the prescription of potentially hundreds of drugs, will challenge the storage capacity required for clinical data. In addition, recent studies have shown the utility of whole genome analysis (6 billion bases of DNA per individual), discovering a cause of Charcot–Marie–Tooth disease, one of the most commonly inherited neurological disorders, which affects approximately 1 in 2500 people. In addition, other disease-causing genes have also recently been found by looking in families that inherit a variety of different types of mutations.
This is no longer the “spit-test” parties and subsequent Single Nucleotide Polymorphism (“SNiP”) analysis that Direct-to-Consumer companies offer to consumers, often only showing modest increased risk for disease prediction (typically ranging from 1.3 – 1.8 fold greater than another individual). Also, as reported here by me in a previous blog, the mutations that lead to Adverse Drug Events often convey a 100 – 10000+ fold greater risk for that patient.
“There will be an explosion of family sequencing that will identify disease genes,” Dr. Leroy Hood, Director of the Institute for Systems Biology, said in a recent interview. “My prediction is that most of us will have our genome sequences done, included as part of our medical records, and it will be an important part of predictive medicine.” This suggests that “all Healthcare IT systems will soon be overwhelmed by patient genomic and pharmacogenomic data.”
As stated by Dr. George Church, Professor of Genetics at Harvard Medical School and Director of the Center for Computational Genetics, said in a Newsweek interview, – The message is not “Here’s your destiny. Get used to it!” Instead, it’s “Here’s your destiny, and you can do something about it!” Diseases result from a combination of genetic vulnerability and lifestyle. If you know you have high risk of certain diseases, it’s in your interest to know and practice the lifestyle that reduces your risk—and the younger, the better.”
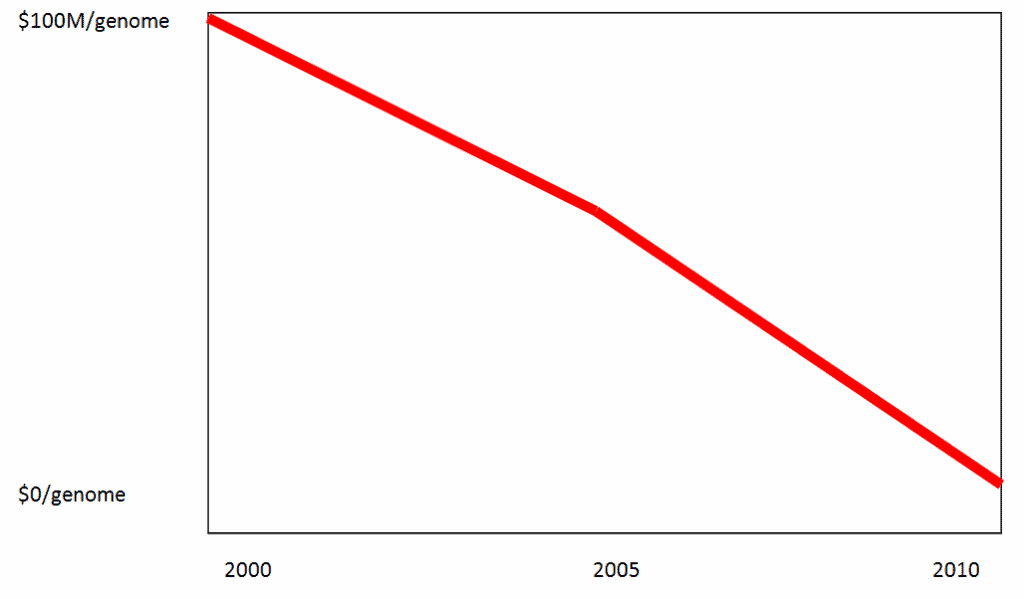
Whole genome sequence analysis, while still expensive, is dropping in price in a dramatic fashion (See Figure below), much faster than the proverbial Moore’s law. The latest research findings have started a “tsunami” in looking at the entire genome in patients with both Mendelein traits and those with common, complex diseases. This has created a need for more computing resources. For example, Washington University’s current scientific and clinical genomic data center, a 16,000-square-foot facility that houses approximately 5,000 processors and more than 5 petabytes of disk storage, is nearly 90 percent full. The University just received a $14M grant from the National Institutes of Health to increase storage capacity.
One obvious solution may be Cloud Computing storage or offsite server farms, connected to the EHR through Web Services by a Secure Socket Layer (SSL), that helps protect patient confidentiality. However, this approach has not fully reached the mainstream of development in the EHR realm.
Adapted from presentation from Dr. George Church at the Cold Spring Harbor Lab, “Personal Genomes” meeting:


marcdparadis says
Gerry – great post, medical genomics and medical proteomics will definitely have significant impacts on the practice of medicine and on quality of life.
A quick postscript, the following companies are working hard to produce affordable whole genome sequencing: Illumina, Complete Genomics, and Pacific Biosciences. AthenaHealth, of course, already has EHR in the cloud and many other EHR vendors will too.
DrByte says
Enjoyed the article: I see three major factors even more critical than the actual storage of genomic data.
1. How will the data become information for the clinician so a decision can be quickly and accurately made?
2. How will the data be displayed so it can be interpreted by the clinician? A long dissertation on a genetic profile will not be appreciated nor correctly interpreted by healthcare providers?
3. What will be the ontology/codification standards for genomics? Without a well-thought out, comprehensive, rapidly responsive and fluid system, the data will be not retrieval, fully utilized or usable.
Gerry Higgins says
DrByte-
You have raised some of the most critical, and completely ignored, components of the need for a usable way for the busy physician to understand and implement genomic data.
In all its arrogance about funding “translational genomics”, the NIH and HHS has put little thought into how to actually present comprehensible genomic data in an EHR or similar HIT application.
Clinical Decision Support Systems may provide one solution – see http://www.warfarindosing.org as an example for accurate determination of Coumidin dose.
I suggest that we need more “intersectionalists” like myself (excuse the grandiosity), who have training in bioinformatics/genomics and who actually work in a non-academic hospital clinical IT setting. There are no good ways to represent genomic data in the EHR, although the NIH has funded some small research projects at places like the University of Virginia (‘The Genome-Enabled EHR’).