An HHS committee on Physician and Patient Education is developing new certification requirements for hospital physicians in genomics and pharmacogenomics. Most hospital physicians, unless your specialty is oncology or obstetrics, are not familiar with anything but diseases inherited as classical Mendelien traits. These new rules will require basic certification in both classical genetics and the “new” genetics of disease and pharamacogenetics of drugs that have been discovered following sequencing of the human genome, and stem from large Genome Wide Association Studies (GWAS) that examine gene-disease and gene-drug associations.
The Secretary’s Advisory Committee on Genetics, Health, and Society has developed a set of recommendations for HHS Secretary Kathleen Sebelius that would provide advice on steps that can be taken to help prepare physicians for dealing with new types of data, and new requests from patients about genetic information, such as reports from direct-to-consumer genomics firms. According to one source, “The task force found that genetics education for health care professionals is inadequate or ineffective in part because of out-of-date formal training and a lack of understanding of the usefulness of genetics in contemporary medicine…The panel also would recommend mechanisms for expanding and enhancing content to prepare health care professionals for personalized genomics, and it would recommend how evolving standards, certification, accreditation, and continuing education activities might be incorporated into genomic content. The panel also would plan to monitor the outcome of its efforts. The task force recommended a systematic effort that evaluates the job responsibilities of the health care workforce in order to target educational efforts that improve genomic or genetic knowledge. It suggested HHS fund the creation of a web-based information resource center that builds on existing government resources, such as those at the National Institutes of Health, that would specifically provide comprehensive, accessible, and trustworthy genetic information for consumers. Physicians and other health care workers, as well as consumers, also should be educated about the importance of using family health histories, the task force said. It advised that HHS support using family histories in clinical care through the development of support tools and mechanisms to integrate pedigrees into electronic medical records.”
At the same time, the FDA is perseverating about how to inform physicians about the dangers of certain drugs. Their usual approach to inform physicians is through the use of the paper-based Drug Information Label (“Learning from Labels”), but many question to what extent prescribers actually read these detailed inserts. At a recent FDA and Drug Information Association (DIA) meeting, the FDA proposed several different ways of informing and educating physicians about the need for gene testing prior to the prescription of certain drugs, including:
Question to Audience: What is the best way to communicate effectively with physicians and healthcare providers about label changes involving a Pharmacogenomic test and its appropriate medical use, particularly for existing drugs?
1.FDA ‘Dear Doctor’ letter
2.FDA web site alerts
3.Drug company representatives
4.Hospital or Pharmacy Benefit Manager
5.Laboratory offering the gene test
6.Medical groups (e.g., American College of Cardiologists)
7.Expert government panels (e.g., USPSTF)
8.Press conference and/or press release
9.Other
The response from the attendees was overwhelming – “Use the Electronic Health Record (EHR), with embedded Pharmacogenomic and Clinical Decision Support as the vehicle for educating and informing the physician in the course of clinical workflow.”
Why is this important? Well for Pharmacogenomics, patients are suffering from side effects and dosing errors that harmful and potentially fatal. Some of the ground-breaking work by Brian Gage, M.D. at Washington University in this field has focused on the development of a dosing algorithm (actually 16 integrated algorithms) that more accurately predicts a starting and maintenance dose for patients taking warfarin, the anti-coagulant. In this instance, genotyping of patients to identify the correct dose is based on genetic variability between humans – using clinical data alone, as might be collected from an EHR, has an accuracy of 12%, while combining clinical data with identification of critical single nucleotide variants (Adverse Drug Response alleles) yields an accuracy of 60%. See www.warfarindosing.org to access this critical Pharmacogenomic Clinical Decision Support application.
There are now an estimated significant 250 alleles that are responsible for Adverse Events in pharmacology, including responsivity based on individual human genetic variation to oncology drugs such as Tamoxifen, cardiovascular drugs such as Plavix®, and psychiatric medications such as Selective Serotinergic Reuptake Inhibitors (SSRIs), and the new anti-psychotic medications such as Seroquel®, which is often used to treat bipolar disease and anxiety disorders.
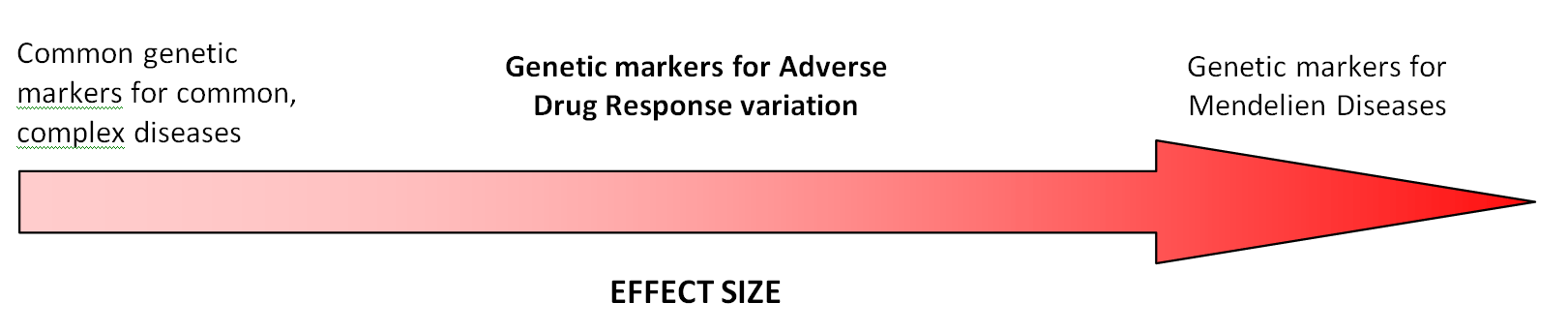
In comparison to the Single Nucleotide Polymorphisms (‘SNipS’) that have shown association with common complex diseases such as Coronary Artery Disease and Major Depressive Disorder, but may only convey an increased risk of between 1.5 – 2.0 fold based on your genomic variation, the Adverse Event alleles often behave more like monogenic traits. Thus, if you have the gene variant(s) that convey an Adverse Event, the risk of having an adverse reaction to a drug may be 10-1000+ times greater than someone without the same genotype. Thus, Adverse Drug Responses fall somewhere between risk for common complex diseases, whose number has grown from 10 known associations to >500 known associations from 2002-2007, and monogenic disease inheritance like Cystic Fibrosis and Huntington’s Disease:
Adapted from Lon Cardon, GlaxoSmithKline:
So what impact will this have on the busy hospital physician? Well, unless you are strongly based in an Academic Research setting, or specialize in domains where genetics are a usual part of clinical care, then you better realize a daunting set of requirements is looming on the horizon.
A solution? Deliver education and information in small amounts in the course of clinical workflow in the EHR. CME, university classes and Dear Doctor letters don’t seem to be the answer, at least in my opinion.



Anthony Guerra says
Thanks Gerry.
What is the relationship between HHS and the private medical schools? Do they look at HHS guidelines as recommendations or are they more forceful, meaning that HHS directives on physician education must be adopted within a certain amount of time?
Most importantly, should CIOs be asking their EHR vendors anything in particular about the ability of their systems to handle genetic information?
Gerry Higgins says
Anthony – As you may know, the Association of American Medical Colleges (AAMC) is the lobbying arm of the U.S. medical schools. They tend to act as intermediaries with HHS, the NIH, the FDA and other federal agencies in situations such as this. Once certification in Medical Genomics becomes mandatory (like ACLS, ATLS), then all physicians will have to obtain certification or will not be allowed to practice.
I don’t believe that the push will come from the medical schools – instead EHR vendors and other commercial entities will fullfill this role.
As I have said in a earlier blog post, there is a great deal of emphasis being placed on the “Genome-Enabled EHR” by HHS – once applications such as Pharmacogenomics Clinical Decision Support becomes implemented, then any prescriber is at risk if they do not follow the results, especially if a patient has an adverse event.
I am telling you, this is like a Frieght Train bearing down on the hospital community, and everyone is asleep at the wheel on the tracks!